The branch of chemical biology based on designing and synthesizing small molecules have provided powerful reagents in forms of enzymatic inhibitors, activity-based probes, and fluorescent reporters to investigate various biological processes. Still, the complexity of the cell is far beyond of the reach of small molecules in our toolbox. We are developing another branch of chemical biology by engineering proteins with new catalytic or molecular recognition capabilities. These designer proteins would provide us new opportunities for discovery at the interface of chemistry and biology. Specifically, we are engineering the E1-E2-E3 enzymatic cascade to map cell signaling pathways based on ubiquitin transfer.
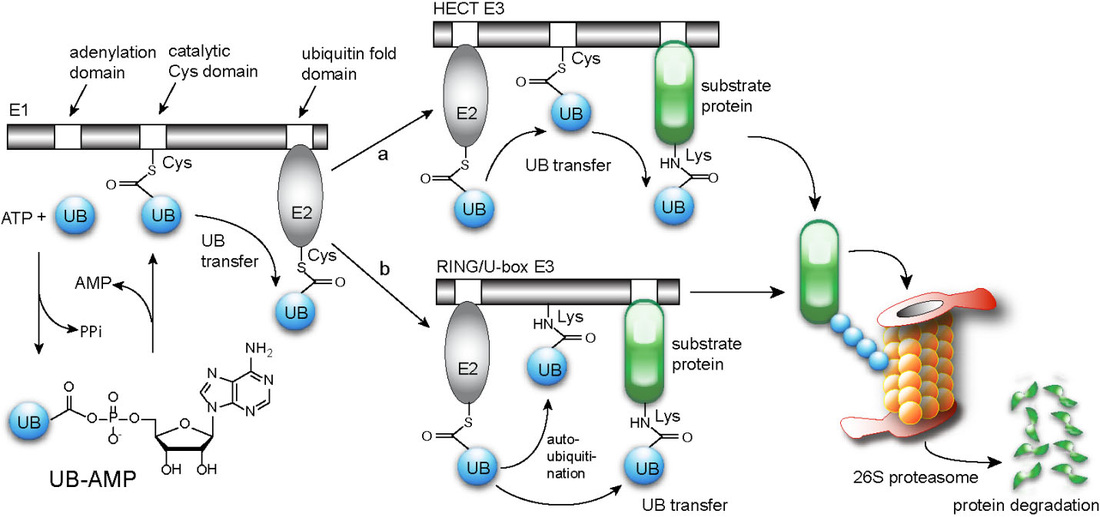
Ubiquitin (UB) is transferred through E1, E2 and E3 enzymes to cellular proteins to regulate their degradation and biological functions (Figure 1). The substantial cross-reactivities of 2 E1s, 50 E2s, and more than 600 E3s make it a significant challenge to identify the substrate proteins of individual E3 enzymes to differentiate their biological functions.
Ubiquitin (UB) is transferred through E1, E2 and E3 enzymes to cellular proteins to regulate their degradation and biological functions (Figure 1). The substantial cross-reactivities of 2 E1s, 50 E2s, and more than 600 E3s make it a significant challenge to identify the substrate proteins of individual E3 enzymes to differentiate their biological functions.
Figure 1. The UB transfer cascade consisting of E1, E2 and E3 enzymes.
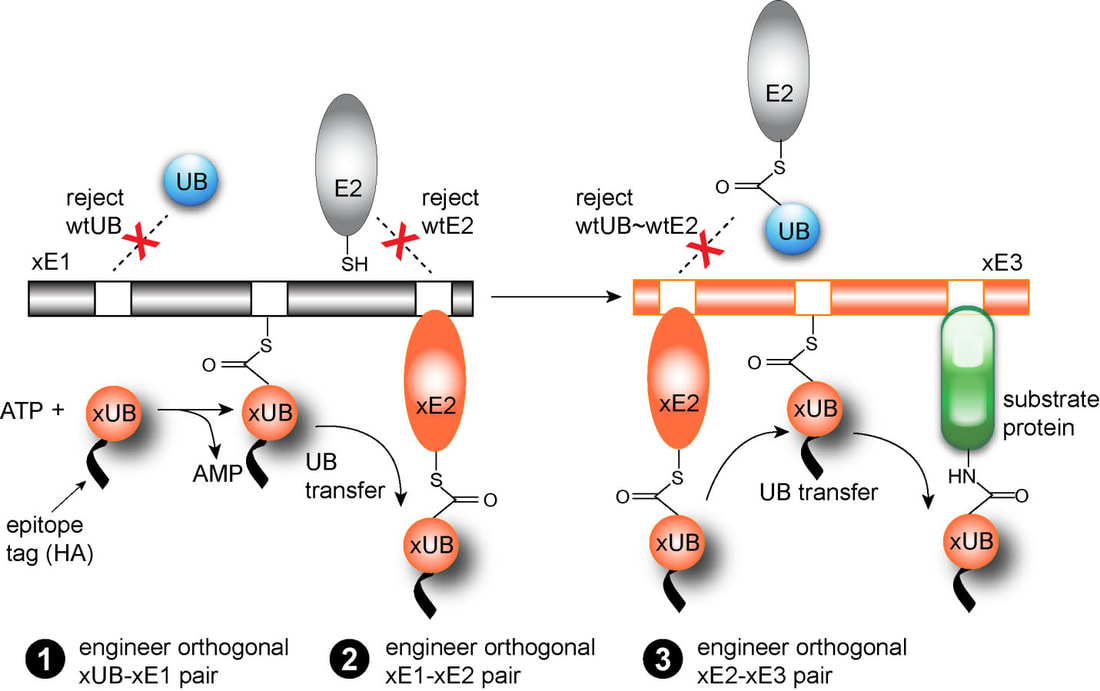
We have developed a method known as “orthogonal UB transfer (OUT)” to profile the substrate specificities of the E3 enzymes (Figure 2). In this method, we engineer orthogonal pairs of xUB-xE1, xE1-xE2, and xE2-xE3 so xUB is exclusively transferred to xE3 and then to its substrate proteins in the cell (“x” designates engineered UB or enzymes). The engineered pairs are free of cross-reactivity with the native pairs so xUB stays in the track of the xE1-xE2-xE3 cascade for delivery to the substrates of a specific xE3 . We call such a cascade for xUB transfer an “OUT cascade”. We express the OUT cascade and xUB fused to 6×His and biotin tag in the cell, and purify cellular proteins conjugate to xUB. We identify these proteins by tandem mass spectrometry (MS), and they are the likely ubiquitination targets of the E3 enzyme. By generating the substrate profiles of individual E3s, we would be able to map various E3s on the cell signaling networks and elucidate their roles in cell regulation.
Figure 2. The OUT transfer cascade and the engineering of orthogonal xUB-xE1, xE1-xE2 and xE2-xE3 pairs.
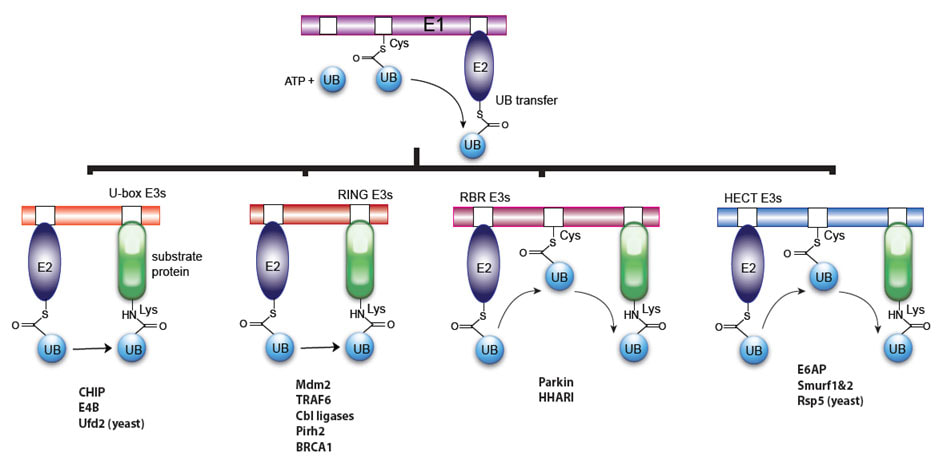
We are now building OUT cascades with various classes of E3s such as HECT E3s E6AP and Rsp5, Ring E3s Cbl-b and Mdm2, Ring-between-Ring (RBR) E3s Parkin and HHARI, and U-box E3s CHIP and E4B. All these E3s play key roles in cell regulation and are implicated in many diseases such as cancer, neurodegeneration, autoimmune, and infectious diseases. By identifying their ubiquitination substrates, we would elucidate their roles in pathogenesis and guide the development of effective therapies.
Figure 3. E3s in the lab for the generation of OUT cascades.